And this looks just....sophisticated enough to be something people would do single cell analysis with
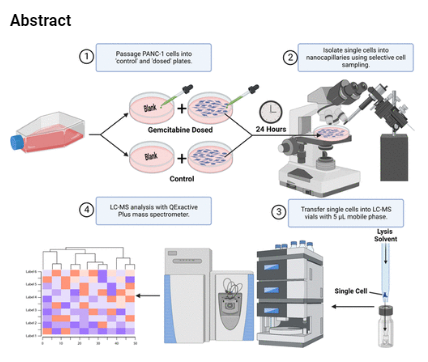
This group used a standard QE Plus operating at 300 microliters per minute (or 350 microliters/minute if using a special column) -- holy crap. I wouldn't have guessed I'd have detected lipids in 1,000 cells. Let alone one at a time. I've got a QE Plus! I'm pretty sure the LC on it can go that high, I generally run it lower so I can have higher sensitivity, though...
Now....apparently the reviewers at ACS did forget to make sure that the RAW files were publicly available, but maybe they don't require it because who cares about lipids?
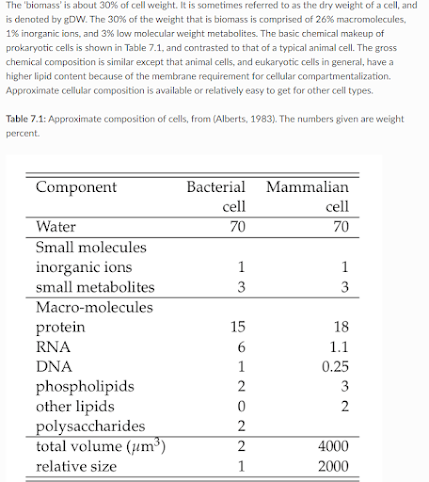
Okay, because I have a rapidly approaching deadline -- what is the lipid content of a single cell anyway? Table 7.1 in this old book (open access here).
5% of the Eukaryote cell content is lipid (probably put a big -ish on this, I can't find the original study of Alberts, 1983 through the references). So....let's go with that's around 50 picograms of lipids per Eukaryote cell. Okay, at least it is more than the 1pg or so of mRNA floating around (about 10% of RNA is mRNA, most of the RNA around is the stable stuff, 16S, etc.,) I'm getting that number off of it being about 1/4 the percent content of protein and a mammalian cell is generally around 200pg (blog post from 2014....ouch...wait, I was still using SpongeBob memes in 2014?)
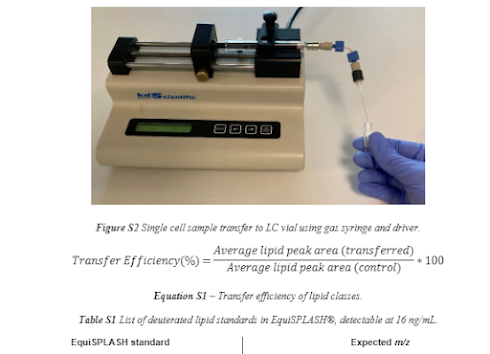
What about their standards? They used the ones we use from EquiSplash and they optimized at 16ng/mL
So, 16 picograms/uL and they put about 5uL injections, so they optimized their method around 80 picograms.
That's not too far off the 50 picograms per single cell, except that's total for phospholipids and other lipids. And since I have no idea what the distribution of different lipid species are (how many and do classes 1-7 make up 90% of them) but -- this actually seems like it is in the right realm order of magnitude or two. Weird, right?

No comments:
Post a Comment