Do you just want to take a mass spectrum from anything - maybe even different instruments or online data or NIST fragments or literally anything else and just make a nice figure?
www.mass-spectrum.com.
Feel free to read the nice tutorial - or just use the most intuitive interface you've ever seen.
YOU CAN EVEN MAKE FIGURES FROM BRUKER DATA! No joke.
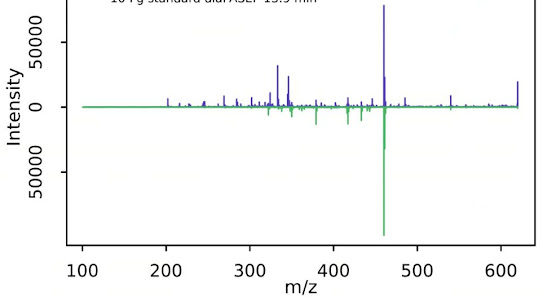
That figure at the very top is me extracting diaPASEF data from a drug standard and from a drug treated complex sample! Bottom is my drug standard.
How do you do this? You get into the hellscape of terribleness that is "Data Analysis(TM)" and you find your spectrum and make it a "Compound Spectrum" then you right click on that spectrum in your table and export it as a peak list .ASC file. Then you open that in Excel and trim out the 3rd column that is just random weirdness, save it as a .text file and load it into www.mass-spectrum.com.
You can add an Excel file with your masses/peak lists and - BOOOM - you've made a labeled plot. Everything from the colors to the fonts to - you name it - is user selectable.
I've successfully made a mirror plot from QE PRM data for a standard small molecule and from a TIMSTOF diaPASEF small molecule run!


No comments:
Post a Comment