I have been buried in work linked to maintaining my employment for several weeks. So buried that I scheduled a lot of blogposts for 2032. I'll leave them there so they'll post again later.
The annual congress of EuPA is something I've always wanted to attend and it has never quite worked out, so I conned some guest bloggers into providing some insight about what happened at the conference. (Sorry for the month long delay).
At Proteomics
Forum/EuPA2022 in Leipzig, we got word of the shout out from across the ocean
to provide a little update on the most exciting stuff that was presented. In
search of a brief format, Ghent attendees (#GhentRepresent) spread out and each
of us came up with a personal perspective on the most exciting stuff, a
predominantly ECR perspective:
Arthur
Declercq: “EuPA2022 was the conference of tackling
problems in specific proteomics niches such as single cell or
immunopeptidomics. It was very nice the see many people showed up for the
immunoproteomics symposium where Michal
Bassani-Sternberg gave her talk about identifying neo-epitopes for
personalized cancer immunotherapy with their proteogenomics pipeline NeoDisc,
showing how they bring state-of-the-art research from bench to bedside (10.1038/s41587-021-01072-6).”
Ben commentary: Hey! This is one of the two immunopeptidomics splicing groups! Despite the controversy about whether HLA peptides splice or not, there are some very successful clinical trials going on right now centered on weird spliced peptides.
Tine Claeys: “The
diversity of proteomics and its wide variety of instrument, tools,
applications, … shows the creativity of scientist in tackling very different
problems. I was intrigued by the talk from Jennifer
Van Eyk on how her group approaches clinical applications and biomarker
discovery integrating multi-omics analyses. Combining already existing clinical
data with the new advancements in the mass spec field to determine your
molecular twin demonstrated again the importance of big-data analysis,
especially in a clinical context. The
talk from Ileana Cristea on
spatiotemporal organelle remodeling ( 10.1016/j.celrep.2020.107943) blew my mind on how the location of a
protein can have a huge impact and can be very altered upon viral infection but
we're still mainly focused on protein abundance. This will definitely impact
the way I approach my own research. I've had some amazing interactions with
amazing people and expanded my scientific horizon in ways that would've been
impossible without Proteomics Forum 2022!”
Toon Callens: “As I
just started my PhD in proteomics (bioinformatics) it was great to see what
everyone is working on and to have a broad overview of the field and the
state-of-the-art. It was also interesting to see overlap with my own projects,
which gave me some new ideas.”
Annelies Bogaert: “Single cells, single cells, single
cells! It is great to see how much progress has been made in this field. Also
great to see how proteomics is contributing to help conquer the covid-19
pandemic. Chris Overall his talk about how they used TAILS to identify
substrates of the SARS-CoV-2 3CLpro protease which gives insights in
how the virus uses this to its benefit, is just one great example (10.1016/j.celrep.2021.109892). While Albert
Heck gave a nice presentation showing that we are evolving to personalized
diagnostic as top down comparisons of IgG antibodies from one person over time
could in the future help diagnose medical conditions (10.1016/j.cels.2021.08.008). Since
I spend some of my time trying to understand better the functionality of
proteoforms (thanks to Neil Kelleher
for introducing this term [10.1038/nmeth.2369]) it was
nice to hear that so many talks also considered them and stressed their
importance as they are often still neglected! And as a last note, this conference
really showed that DIA is becoming the standard over DDA.”
Tim Van Den Bossche:
“I was really honored to present the flagship manuscript of the Metaproteomics
Initiative and give a short introduction about the Initiative itself. Where the
field 10 years ago was often overlooked, it's amazing to see that there's now a
dedicated metaproteomics session at the major European proteomics conference”
Ben commentary: This is the metaproteomics initiative mentioned: https://metaproteomics.org/
Ralf Gabriels: “Proteomics
is done with simple setups; more challenging workflows is where the interest is now: Single cell, subcellular/spatial, immunopeptidomics, metaproteomics,
proteogenomics, open modification... And quantification labels are coming to
DIA!? Both non-isobaric with plexDIA (10.1101/2021.11.03.467007) and isobaric with DIA-TMT (10.1016/j.mcpro.2021.100177).”
Ben commentary:
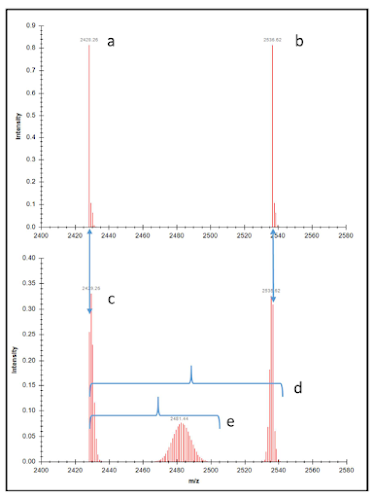
Maarten Dhaenens: “plexDIA,
using 3-plex nonisobaric mass tags: how could I have missed this one? (Nikolai
Slavov) (10.1101/2021.11.03.467007).
Neil Kelleher is doing stuff with single ions
I will never understand because I am a QTOF person, but he is so right that “we
should measure what we must, not what we can!”.
Markus Ralsner suggested
ordering your differential proteins according to chromosome localization,
because you might in fact be looking at an aneuploidy event in your cell lines.
That reminds me of a shrimp project where we once were thinking along the same
lines, but we never really showed it as beautifully as Markus did!
And OMG, Kathryn
Lilley is leaving us to look at nucleotide biomolecules?! Nooooooooo! But
then again: what she is noticing on subcellular RNA localization is beyond
cool. Please Kathryn, come back when you finished throwing over their world
view and throw over ours a few more times?
Vadim Demichev got
across the message that Match Between Runs (MBR) is making a significant difference
in the current DIA-NN implementation. Therefore, he does upfront peak picking
and alignment. And hey, that reminded me of something we were doing a while
ago...
In fact, just talking about it became the biggest breakthrough for me
personally: our ion-networks could finally be revived (10.1101/726273), all because of the return of in-person interactions
and how these are so different from digital meetings! I am convinced that
Bruker data will turn out to be the perfect match for ion-networks. So, if
ion-networks are a “sleeping beauty”, then Proteomics Forum/EuPA2022 sure woke
her up!”
Ben commentary: whoa....check out the highlighted stuff....this happens all the time in cancer, particularly when cells are treated with platinum based therapies. Worth considering, for real...
Lennart Martens: “Lots
of great interactions and creative new ideas at the first in-person meeting in
ages. Also… CAKE!”
Ben commentary: Wait. THE Lennart Martens? Holy shit... This was a great idea!
CONCLUSION: The
conference hall was part of the Zoo of Leipzig. After the conference, we
visited the zoo and found a fish that perfectly illustrated how our heads felt
after a full week of in-person conference!
Guest reporters:
Arthur Declercq (@DeclercqArthur) 1,3
Tine Claeys (@TineClaeys1) 1,3
Toon Callens (@ToonCallens) 1,3
Annelies Bogaert (@AnneliesBogaert) 2,3
Tim Van Den Bossche (@tvdbossche) 1,3
Ralf Gabriels (@RalfGabriels) 1,3
Maarten Dhaenens (@MaartenDhaenens) 4
Lennart Martens (@CompOmics) 1,3
1. CompOmics,
VIB-UGent Center for Biotechnology, VIB, Ghent, Belgium
2. Gevaert Lab, VIB-UGent Center for Biotechnology, VIB, Ghent, Belgium
3. Department of Biomolecular Medicine, Ghent University, Belgium
4. ProGenTomics, Department of Pharmaceutics, Ghent University, Belgium