In advance I want to apologize if this post comes off soundy a little sarcastic. That isn't my intention. We all know that sometimes people who do great science are not good at communicating said science. One example might be the MSStats thing that most mass spectrometrists have heard of but is only accessible to people that commonly compile code.
MSstatsShiny has a pretty web interface but you can only use it if you have a small input file and only if you know the absolutely top secret - you can spend 3 days looking for it and you won't find it anywhere - annotation file format!
If you want to run it locally, all the steps are at the bottom of this post and you still need that annotation file format.
I broke out the big guns here and had one of the best real bioinformaticians on the planet wade through pages and pages of stuff that looks like this
Want it? Too bad, I didn't spend 3 days looking for it AND cash in a huge favor just to give it away for free.
J/K, but y'all owe me. Download it here.
With that out of the way, let's go through THE MASS SPECTROMETRISTS GUIDE TO INSTALLING AND RUNNING MSstatsShiny!
First you're going to need a pain in the butt called RStudio, and you'll need the newest one. You can get it here.
Once you install it, you'll want to open it and go to the console tab.
Now you're going to need bioconductor. You can get that here: https://www.bioconductor.org/install/
Bioconductor is a big package that is necessary for installing all sorts of other things. It's like how you have to install Steam to get games from the Steam store.
Anytime you see something in a grey box on Bioconductor like this, you can just copy and paste this thing and then you put it in the R console by the "greater than" sign. In this context it is probably called a carrot.
Protip for R. If your console has been doing stuff and you don't see a greater than sign, go to the top and hit the little stop sign. Chances are it is running something in the background.
Copy and paste the thing in the grey box to the lowest greater than sign on your screen and hit enter.
Now, I don't know why, but when you are installing stuff IT WILL LOOK LIKE YOU JUST TRIGGERED A NUCLEAR WAR FROM YOUR COMPUTER. You'll see what I mean. The angriest red scrolling text that you've ever seen, generally means things are going okay. Some nerd in Australia found it funny and no one has ever changed it.
Protip for R #2: What I've found with the catastrophe that Windows has become is that my program files will sometimes be locked as read only. You may have to follow the directions in the R console to the file it can't write to then right click, get the little box to pop up, go to properties and unclick "read only".
Once the stress of what a terrible thing you've obviously done is over. It's time to do it again!
Go get MSStatsShiny here! https://bioconductor.org/packages/release/bioc/html/MSstatsShiny.html
Copy the stuff in the grey box to your greater than sign and avert your eyes from the terror on your screen!
It might ask you questions about things you don't know about while you're installing. Just say yes. It will probably be fine.
NOW IF YOU GET THROUGH THIS HERE IS THE NEXT SECRET.
To run the program you'll need this top secret code: (I read somewhere that I might not need all these words, but this is what works for me)


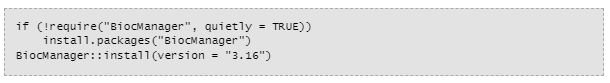

No comments:
Post a Comment