One of the reasons that proteomics has started to take center stage over genomics is the increasing realization that some of the most terrifying diseases aren't really genomic in nature.
Germ line mutations (the stuff you're born with, thank you ancestors!) and cancers are diseases that are due to genetic changes. Deleterious cancer mutations propagate because a mutation in a cell give it a selective advantage over the cells around it so it out-divides them. What about cells that don't divide and are not genomic in nature? You can spend ten million dollars getting read after read after read and not learn a darned thing except maybe invent a new harder way to not learn anything.
Neurons don't divide. DNA alterations aren't propagating
Cardiac cells don't divide. DNA alterations aren't propagating.
While you could argue that all diseases are protein level diseases, these are kind of home runs, right?
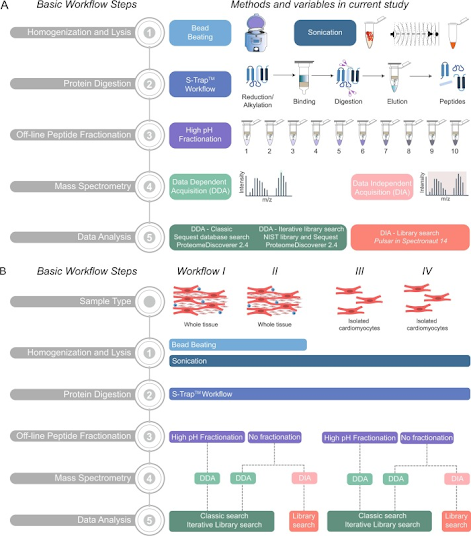
Unfortunately both are a complete pain in the ass to work with. What if you wanted to dive into the latter? Would an incredibly well detailed how-to guide that someone else optimized help? Can I get hell yeah from the back? Thank you!
If you don't take anything away from this great optimization study before jumping right in on cardiac proteomics and following these steps, keep these 2 sentences in mind....
...it sounds like that thing in your chest doesn't beat every second of every day for your entire life because it is simple but if we want to explore it's secrets this tells you where to start!

No comments:
Post a Comment