Just north of Baltimore there is a surprisingly large population of Amish people. You can tell when they're around up here because people will tailgate the buggies and blast their car horns at them. It is likely because people are in a hurry, but I bet some percentage is just tired of all the animal abuse stories that are directly linked to a 40,000 person Pennsylvania population.
What is interesting from a scientific perspective is that this population really hasn't moved about much over the last century or so and there is a lot of genetic data out there on them thanks to things like Geissinger (sp?) which has kept this information.
What can you do with that information? Maybe you could explore functionally enriched genetic variants and then figure out what those gene products do with proteomics?
Sure you could. But why would you want to? Well...what if this population also had a phenotype that a lot of other people would like to have? Like really low levels of the bad cholestorol (LDL)?
And they'd strike out.
Because, while there are protein level alterations in this population (which these authors have previously explored and got one word "Science" for their efforts -- they summarize protein level changes in the MCP paper we're actually talking about) this isn't the most interesting aspects -- or the ones easiest to link to the observed phenotype in question.
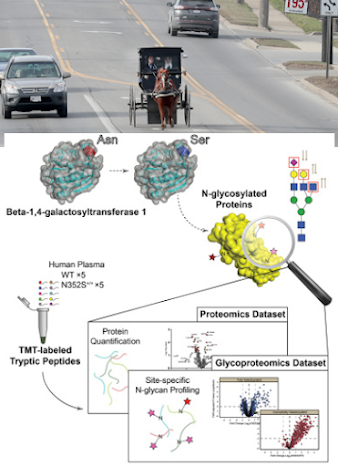
Because (I heard you can't start sentences with because so I'm going to do it multiple times just to prove you can) what appeares to be happening is that the product of this missense mutation is altering the glycan profiles on what the authors describe as "critical targets".
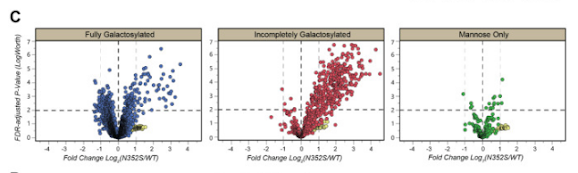
Check out the middle panel in this stolen screenshot:
That looks pretty striking, right?
So you've got this missense mutation enriched in a small population that can improve human health. Ain't a genetics tool on the planet or a protein microarray panel or aptamer or other global panel out there that can tell you how this is functioning, but you can find it through the flexibility of a mass spectrometer and some people who know how to make it do tricks.
This was TMT labeled plasma proteomics analyzed on a QE HF-X. The identifications were made by Byonic from Protein Metrics. Cool story, right?

No comments:
Post a Comment