If LCMS proteomics is going to scale up to the levels of new proteomics technologies that are willing to compromise specificity for throughput, we need to trim down the 4,512 different sample prep methods each lab has sitting around in stacks. You can't tell make a sensible argument that about 4,400 of them are unnecessary.
What we need is some people to do the most boring studies imaginable just to get them out of the way.
OMG. From the pit of my heart, I can't thank these authors enough for this study. It should be referenced by everyone in proteomics all the time. "What do you think about method A vs method B"? I have a hunch but nothing robustly tested but ---
-- these awesome people tested them all! (Well...not all 4,512....but a whole ton of them!)
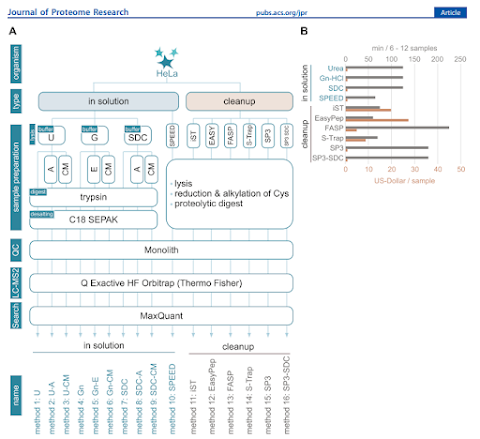
Sure -- number of proteins are one metric -- but they ALSO break down the relative costs of the different methods! Proteomics sample prep has historically been pretty cheap from a reagent perspective compared to DNA/RNA assays that still require hundreds of dollars in reagents. But when you're setting up a 100 sample or a 4,000 sample study, those filters or traps and C-18 spin columns (and TRYPSIN) costs don't seem that irrelevant anymore. On a big study I recently realized that the 1:10 trypsin ratios recommended for some commercial kits was going to blow through every vial in our -20C in very short order. This group considered these factors!
What wins? Nah, you gotta read this so you see how awesome it is. And you've got to remember to cite it later!

No comments:
Post a Comment