Okay y'all, get this. Those trap columns that you just use for making sure your collaborator's samples don't ruin your nice instrument are actually just normal columns that are smaller! Not kidding. You probably wouldn't have realized this from the fact that they say misleading things on them like what resin is inside them and length that they are.
This royal chemistry society advance exploits this fact!
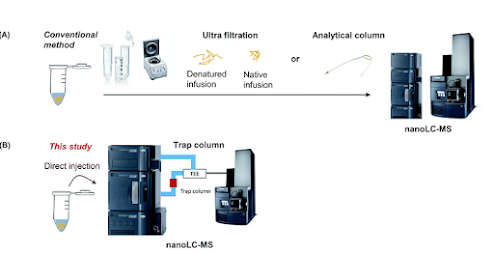
In this advance, this team skips the desalting steps for intact protein analysis and -- get this -- they run their protein over a column while allowing the first part of the liquid that goes through the column TO GO TO WASTE! The salts from their intact protein don't go into the instrument. It goes to some waste thing. Then they elute off of that trap column as if it's a short little column. And the proteins come off of that column and into the mass spec saving untold amounts of time.
I largely like this paper because I think that it you could potentially read my impression of it in two different ways. 1) If you didn't know that you could do this, legit, bravo to these researchers and this nice data. Not kidding. Not being sarcastic. I will be citing this paper all the time going forward.
2) If you've been doing intact protein analysis like this for the last 10 or 20 years and are thinking -- obviously?? OMG, WTactualF?!?!? this should impress upon you how poor we are as a field of passing our information on in a structured and universal matter.
Mass spectrometry is still more like an art than a science in a lot of ways and that isn't a good thing at all. Maybe there is a paper that shows this would totally work out there? I can't think of one and I couldn't find a paper with a Scholar search this morning.
I'm absolutely going to cite the shit out this paper.

No comments:
Post a Comment