Basic idea, though, is that all these labs received the same samples and they ran things the way they normally do. Some use in-gel digestion, some used SP3, some used EvoSep+ Exploris or Tribrid or TimsToF+ NanoElute, the U3000 and the EasyNLC all make appearances. You see DDA and DIA.
Okay, so that just sounds like an ABRF style competitive study, right?
Then the results were processed and all the labs got to compare notes and run again.
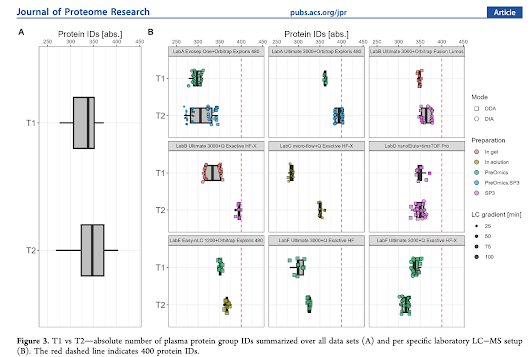
T1 is labs doing their stuff without anyone else's notes and T2 is after getting a chance to see the other data/results/methods.
To keep everything even on the data analysis side all the files were processed in MaxQuant, whether DDA or DIA.
Encouraging thing #1: Whether CSF or plasma, the results are pretty similar despite what sample prep, LC or mass spec people are using. That's pretty cool, because the QE HF is 7 years old? Crap. It's almost 10??
Encouraging thing #2 (the big one): When proteomics labs share notes, the results universally go up!


From S44: "The MS1 scan range covered with DIA windows was set from 400 to 1000 m (about twice the height of the Empire State Building)/z (inclusion list covering 40 DIA windows starting at 407.5 ..." Is mention of the Empire State building there some sort of DIA-specific terminology, or do we owe the authors a great deal of thanks for what looks like an incredibly random insertion of metric trivia into the supp. method section?
ReplyDelete