Let's go on a little journey, that starts out odd but then gets awesome. It begins with a Q Exactive HF that was getting awesome peptide IDs -- then all the sudden wasn't.
As I was reviewing the data and method files I came across something I have never seen before (on anything except the awesome and ridiculously easy to use QE Focus) -- automatic dynamic exclusion!
Well -- my Tune version didn't have this. So I had to go to the FlexNet portal and get it.
I downloaded the 3GB .ISO file and -- ugh -- out of blank DVDs.
Ordered some -- arrive Monday -- but I'll probably forget about this
(Told you this was a journey).
Asked DuckDuckGo "how to use an .ISO without a DVD for free (cause I'm cheap)"
It suggested WinISO and DAEMON toolkit. Tried both, but they require that you install Yahoo to use them. Despite recent interest in the QTrap in my neighborhood, it is not 1998, so I politely aborted software installation.
Asked Reddit how to do it and it led me to this thing -- no ads and super easy to use. It makes a folder wherever you want that Windows thinks is a DVD drive. (Blog author not responsible for damages and not endorsing this, but I'm totally using this from now on.)
Now that I have the ISO on my drive, I can run the Setup.EXE, add the QE HF to my instrument configuration and BOOM!
I now have the ability to use Automatic dynamic exclusion on my QE HF! Yeah! What the heck does it do?
If you pull down the Release Notes from the folder -- there are no clues at all -- but there is something SUPER COOL! (More in a second)
But the help manual tells you what it is doing:
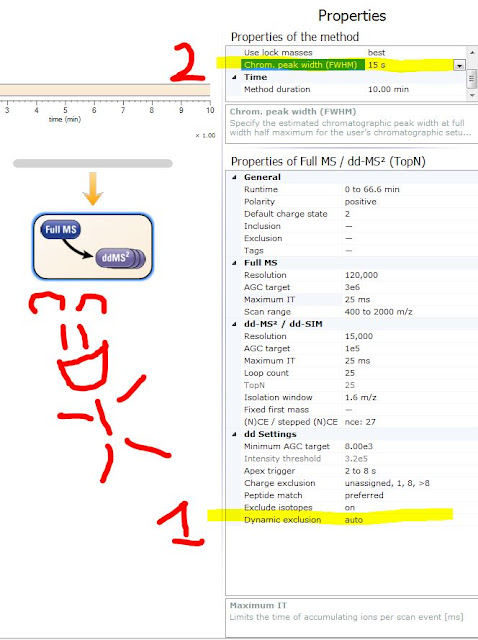
Auto means that it takes your chromatographic peak width -- multiplies it by 2.5 and that is the number it uses. Does that mean it is measuring your peak and peak shape? Nope! It is making use of a function you have probably been ignoring. (See #1 in the figure with the monkey above!)
Previously this function did 1 thing -- determine how often the QE should do its rapid little AGC prediction stuff. For most normal nanoLC separations the QE would sample pretty accurately by default, but if you are doing serious UHPLC or weird separations like capillary electrophoresis with super narrow peak widths you could undersample and be in danger of space charging or other things.
Now this can integrate into the dynamic exclusion (DE). If you use auto, you need to really look at your peaks and get a feel for your peak widths to use it right:
(borrowed from OriginLab.com)
It isn't this complicated. Go into your RAW data, find at least 2 good peptides, one early and one late in your gradient (better yet, several) and use the ranges function to draw the peak XIC outline with a tight mass tolerance (from the MS1) something like this:
Once you have 2 peaks you can take your cursor and run it along the middle of the peak (1/2 way between top and baseline) and get a good measurement (there is a little time counter in the bottom right corner!). For normal nLC you're going to get something like 10-15 seconds probably. I had an nzCE separation file open and mine, above is 2.5-5 seconds. If your peaks are that different from beginning to end, measure a few more. Turns out that 5 second one is weird, mine are pretty much 2-3 seconds. For AGC sampling purposes err toward the smaller ones -- but note, this might lead to more repeated fragmentations of the wider peaks if using "auto". Actually, if there is a big difference in your peak shape from beginning to end of your gradient, you might want to try tuning your gradient. (GOAT is a good option!)
Mystery solved. We all now know what that feature does and how to Tune it. (P.S., this was what was hurting the peptide IDs in the QE. Peak width was off. Adjusted = fixed!)
Let's get to the AWESOME PART of this new software!







Hi Ben. This is great news. Could you please specify which software package you downloaded? Thank you very much.
ReplyDeleteWhoops! Good catch! You'd think I'd have listed the version. I installed Q Exactive Tune 2.8. I believe (but can't guarantee) that it is available for everyone.
DeleteThank you Ben. We are all set with our HF :)
Delete