Did you know that there are specific breast cancer cell lines that are known to preferentially metastasize to the brain? I didn't! They have one called MDA-MB-231BR (please excuse typos) that has this terrifying behavior (they actually call it brain-seeking behavior...
...sorry...had to lighten this up some..)
They do membrane enrichment by ultracentrifugation at 150,000 x g and quantify proteins in this cell line and other cells from the same tissue that aren't known to have this trait -- AND they compare the quantitative membrane profile to a brain cancer cell line. The proteomics is single shot in triplicate using a standard "high/low" method on an LTQ Orbitrap Velos system. (They also spend some time optimizing their membrane prep procedure).
Data processing is done in PD 1.2 -- and quan is done with spectral counts in Scaffold.
The general idea is that there might be some sort of pattern at the cell membrane that is up-regulated in the brain seeking line, not present in the others, that would confirm in the brain cancer line.
Using their method, they find a series of potential candidates.
I downloaded this data from ProteomeXchange here (PXD005719). I reprocessed the data with SequestHT/Percolator/Uniprot-SwissProt and get very similar protein numbers to what they find.
This brings me around to why I was downloading their data anyway. I was curious to see how a membrane prep proteomics procedure compares in coverage to an ultradeep proteome such as the Bekker-Jensen et al., dataset.
I pulled the breast cancer cell line from the Bekker-Jensen repository and processed it side by side with the MB-231-R RAW files from this study. There is some apple-to-oranges going on here, vastly different chromatography, different instruments, etc., but a comparison at the protein level is pretty interesting (at least to me). The overlap at the protein group level (thanks Venny!) looks something like this:
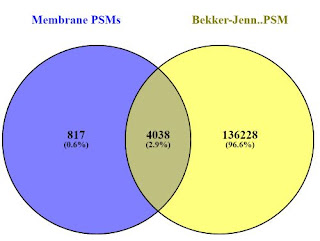
And if we compare the PSMs (P.S. Chrome does NOT like when you load 390,000 peptide sequences into it. It kept throwing me popup errors, but eventually completed. Also worth noting, Venny removes duplicates so the PSMs actually match the peptide group counts.
Honestly, I was a little surprised and checked this again. Considering the absolutely amazing depth achieved in the Bekker-Jensen experiments, I did not expect that 10% of the proteins identified by Peng et al., to be novel to their set. Sure, it's less than 80 proteins, but it appears to be at least a couple of the ones downstream analysis tells them are interesting.
This obviously isn't a perfect experiment. Honestly, this is pretty bad science. It's completely possible that the 76 proteins found in the MD-231-R aren't even expressed in the cell line used in the other study. I could eliminate factors like parsimony and grouping errors with a better experimental design on my side, and may still do that (Bekker-Jensen cell lines take 8 hours on my PC -- without LFQ...so I'm not in huge rush...) I'm putting this here because I came into this little experiment with a preconceived expectation of the outcome, that perhaps if the proteomic analysis was deep enough that membrane enrichment isn't as necessary as we once thought it was. I'm going to leave this lazy analysis here because it seems to suggest the opposite.
Oh -- and this is a good paper with an interesting experimental design you should check out!
(Dorte, if you happen to see this, sorry about the extra "n"s all over the place.)



No comments:
Post a Comment